fig5
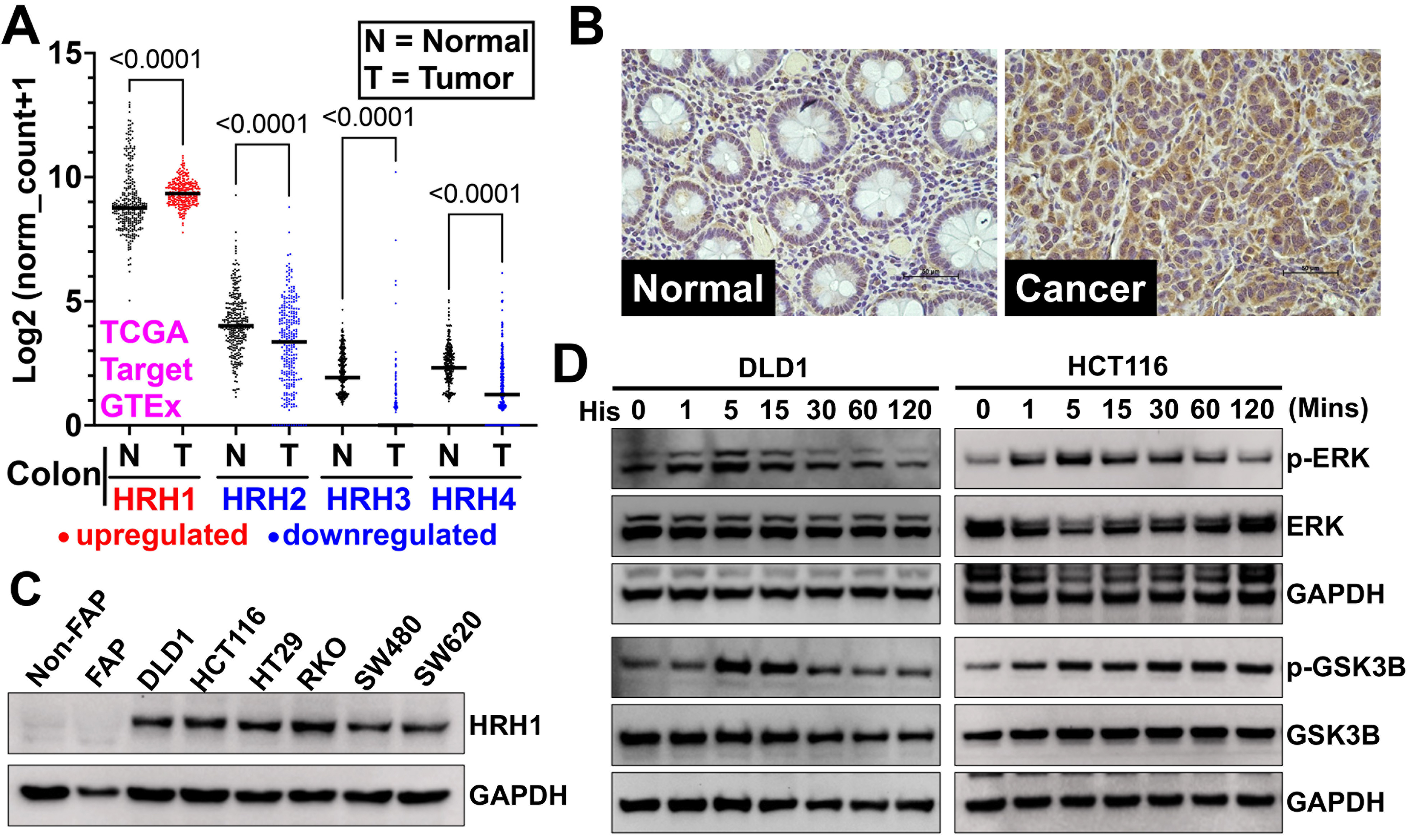
Figure 5. HRH1 is overexpressed and functional in CRC tissues and cell lines. (A) We analyzed HRH1-HRH4 gene expression in colon cancer patient tissues from the TCGA and TARGET databases using the Xena browser. We then compared it to adjacent or normal colon tissues from the TCGA and GTEx databases. HRH1 expression was significantly upregulated, while HRH2-H4 expression was significantly downregulated in colon cancer tissues compared to adjacent normal colon tissues. Ordinary one-way ANOVA was used for statistical analysis; (B) Immunohistochemistry of a CRC tumor microarray (adjacent Normal = 15, colon adenocarcinoma = 15 tissues), obtained from Tissue microarray cat no. CO243b and BRCF of KUMC showed overexpression of HRH1 in colon cancer compared to its adjacent normal colonic tissue (scale bar = 50 µm); (C) Western blot analysis revealed that HRH1 was upregulated in CRC cell lines compared to non-cancerous colonic cells. HCT116 and DLD1 were selected for further study due to their high expression levels among the tested CRC cell lines; (D) Western blot of lysates from CRC cells, previously synchronized in serum-free media for 24 h, treated with 10 µM of histamine for up to 2 h in serum-free media, showed a time-dependent increase in ERK1/2 (Thr202/Tyr204) and GSK3B (Ser-9) phosphorylation. HRH1: Histamine receptor H1; CRC: colorectal cancer; HRH4: Histamine receptor H4; ANOVA: analysis of variance; BRCF: Biospecimen Repository Core Facility; KUMC: University of Kansas Medical Center; ERK1/2: extracellular signal-regulated kinase 1/2; GSK3B: glycogen synthase kinase 3 beta; p-ERK: phosphorylated-extracellular signal-regulated kinase; p-GSK3B: phosphorylated-glycogen synthase kinase 3 beta; GAPDH: glyceraldehyde-3-phosphate dehydrogenase.