fig5
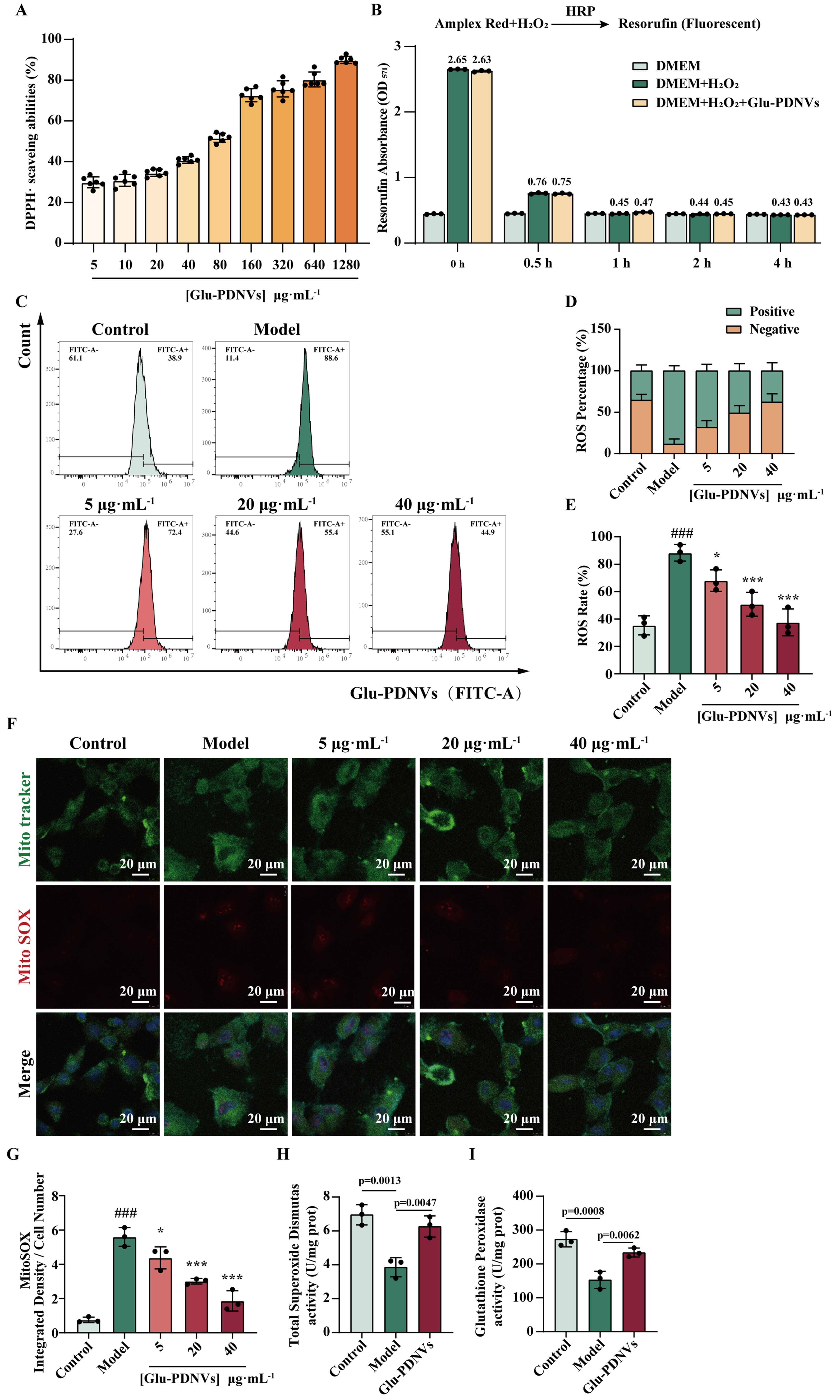
Figure 5. Glu-PDNVs effectively protect H9c2 cells from oxidative damage caused by H2O2. (A) The inhibition rate of Glu-PDNVs against DPPH· radicals at gradient concentrations. n = 6; (B) Direct H2O2 scavenging capacity of Glu-PDNVs. n = 3; (C) ROS levels in H9c2 cells were quantified by flow cytometry; (D) Quantitative analysis of ROS-positive and -negative cell populations; (E) Representative flow cytometry histograms of intracellular ROS levels. n = 3; (F) Representative fluorescence images and (G) statistical analysis of MitoTracker and MitoSOX staining in H9c2 cells. n = 3; Effects of Glu-PDNVs on endogenous antioxidant enzyme activities in H2O2-injured H9c2 cells, showing (H) SOD activity and (I) GPx activity. n = 3. Data are presented as mean ± SD and analyzed by one-way ANOVA (###P < 0.001 vs. control; *P < 0.05, ***P < 0.001 vs. model). PDNVs: Plant-derived nanovesicles; Glu-PDNVs: Rehmannia glutinosa-derived nanovesicles; H2O2: hydrogen peroxide; HRP: horseradish peroxidase; Amplex Red: 10-acetyl-3,7-dihydroxyphenoxazine; DMEM: Dulbecco’s Modified Eagle Medium; DPPH·: 2,2-diphenyl-1-picrylhydrazyl radical; ROS: reactive oxygen species; FITC: fluorescein isothiocyanate; MitoTracker: mitochondrial tracker dye; MitoSOX: mitochondrial superoxide indicator; SOD: superoxide dismutase; GPx: glutathione peroxidase; OD: optical density; ANOVA: analysis of variance; SD: standard deviation; µg: microgram; mL: milliliter; µm: micrometer.