fig1
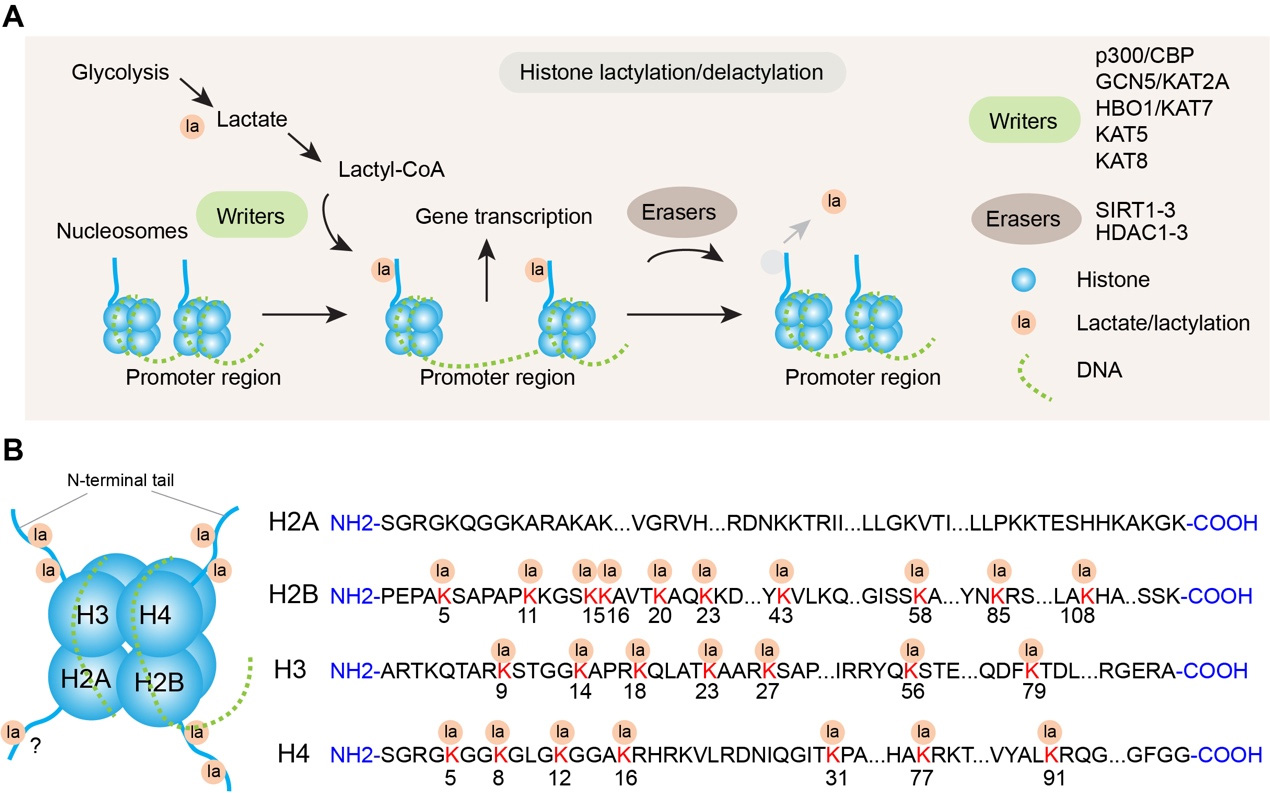
Figure 1. Histone lactylation as a metabolic-epigenetic regulator. (A) Schematic of histone lactylation-mediated gene transcription regulation. Glycolysis generates lactate, which is converted to lactyl-CoA, a key substrate for histone lactylation. “Writers” (e.g., p300/CBP, GCN5/KAT2A) catalyze the addition of lactyl groups (la) to histones in the promoter region of nucleosomes, leading to the activation of gene transcription. Conversely, “Erasers” (e.g., SIRT1-3, HDAC1-3) mediate histone delactylation, removing lactyl modifications and reversing transcriptional activation; (B) Identified lactylation sites on histone N-terminal tails. Lactyl groups (la) are enriched on the N-terminal tails of core histones (H2A, H2B, H3, H4). The sequence of each histone is shown, with numbered positions (e.g., H2BK5, H3K9) indicating confirmed lactylation sites. Note: modification sites in H2A have not been identified in the canonical H2A sequence, despite a previous report[11].